Single-cell, single-nucleus and xenium-based spatial transcriptomics analyses reveal inflammatory activation and altered cell interactions in the hippocampus in mice with temporal lobe epilepsy
单细胞、单核和基于氙的空间转录组学分析揭示颞叶癫痫小鼠海马中的炎症激活和细胞相互作用改变
| 期刊: | Biomarker Research | 影响因子: | 9.500 |
| 时间: | 2024 | 起止号: | 2024 Sep 13;12(1):103. |
| doi: | 10.1186/s40364-024-00636-3 | 研究方向: | 神经 |
| 疾病类型: | 癫痫 | 细胞类型: | 其它细胞 |
文献解析
1. 领域背景与文献引入
文献英文标题:Single-cell, single-nucleus and xenium-based spatial transcriptomics analyses reveal inflammatory activation and altered cell interactions in the hippocampus in mice with temporal lobe epilepsy;发表期刊:Biomarker Research;影响因子:未公开;研究领域:颞叶癫痫(TLE)的海马病理机制研究。
颞叶癫痫是临床最常见的局灶性癫痫类型,约占所有癫痫病例的40%~60%,患者常因反复癫痫发作出现认知障碍、情绪异常及精神症状。其核心病理改变为海马硬化(HS),主要表现为海马神经元进行性丢失、胶质细胞增生及神经纤维密度增加,但海马硬化的发生机制及癫痫的起始与进展过程仍不明确。近年来,神经炎症被认为是癫痫发生的关键驱动因素——小胶质细胞激活(如CD68表达增加)和星形胶质细胞反应性增生(如GFAP上调)可通过释放促炎因子(如IL-1β、TNF-α)加重神经元损伤;同时,神经元的兴奋性-抑制性平衡破坏(如GABA能神经元丢失)是癫痫发作的核心机制。然而,现有研究多基于bulk转录组测序或单一单细胞技术(如单细胞RNA测序难以有效捕获神经元,单细胞核RNA测序对胶质细胞覆盖不足),难以区分细胞类型特异性的转录变化;且缺乏空间分辨率的研究无法解析海马内细胞的空间分布及互作,限制了对癫痫发生机制的深入理解。针对这一研究空白,本研究整合单细胞RNA测序(ScRNA-seq)、单细胞核RNA测序(SnRNA-seq)及Xenium空间转录组学技术,对kainic acid(KA)诱导的颞叶癫痫小鼠海马进行分析,旨在构建高分辨率的细胞类型特异性转录组图谱,揭示海马炎症激活与细胞互作改变的分子机制,为海马硬化及癫痫发生的早期干预提供理论依据。
2. 文献综述解析
文献综述以“技术局限性-现有结论-未解决问题”为核心逻辑,对颞叶癫痫的研究现状进行系统评述。现有研究已明确:①海马硬化是颞叶癫痫的特征性病理改变,与神经元丢失、胶质增生及神经纤维重构密切相关;②神经炎症是癫痫发生的重要驱动因素,小胶质细胞激活和星形胶质细胞反应性增生可通过促炎因子释放加重神经元损伤;③神经元的兴奋性失衡(如谷氨酸能神经元过度激活、GABA能神经元丢失)是癫痫发作的核心机制。现有研究的优势在于识别了一系列与炎症和神经传递相关的分子标志物(如Spp1、BDNF),为理解癫痫发生提供了线索;但局限性同样显著:bulk测序无法区分细胞类型,单一单细胞技术难以同时覆盖胶质细胞和神经元,且缺乏空间信息无法解析细胞互作。
本研究的创新之处在于突破技术局限,通过三种测序技术的整合实现“细胞类型+空间维度”的双重解析:①利用单细胞RNA测序解析胶质细胞转录组,单细胞核RNA测序解析神经元转录组,弥补了单一技术的不足;②通过Xenium空间转录组学获得基因的空间分布信息,揭示胶质细胞和神经元变化的空间特异性;③在癫痫发生的潜伏期(KA注射后7天)进行研究,捕捉到海马硬化和癫痫发生的早期分子事件。研究结果不仅验证了现有研究中炎症通路激活的结论,更识别出跨胶质细胞类型和空间数据一致上调的分子(如Spp1、Trem2、Cd68)及神经元中的特异性分子(如Penk、Sorcs3、Plekha2),为颞叶癫痫的生物标志物开发和治疗靶点筛选提供了新的候选分子。
3. 研究思路总结与详细解析
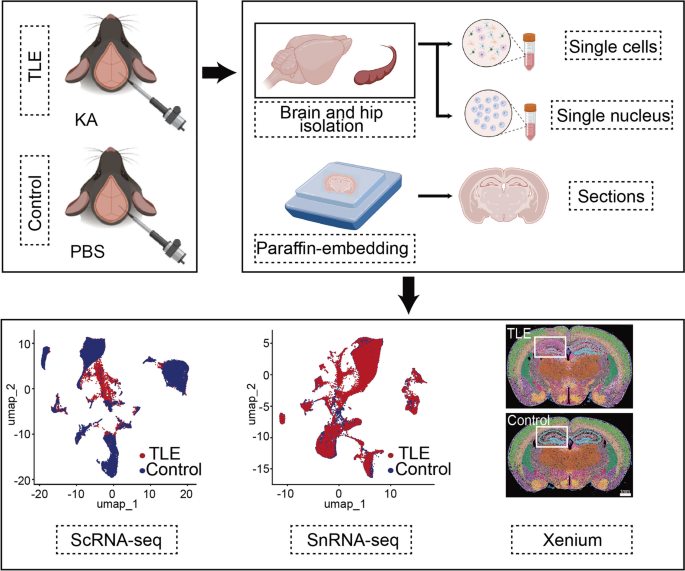
本研究整体遵循“模型构建-多组学测序-生物信息学分析-结果整合”的闭环思路:首先通过KA立体定向注射构建颞叶癫痫小鼠模型,然后采用单细胞RNA测序(解析胶质细胞)、单细胞核RNA测序(解析神经元)及Xenium空间转录组学(解析时空分布)对海马样本进行测序,接着通过生物信息学分析鉴定差异基因、功能通路及细胞互作,最后整合多组学数据揭示炎症激活与细胞互作改变的机制。
3.1 颞叶癫痫小鼠模型构建
实验目的是诱导具有海马硬化特征的颞叶癫痫模型,为后续研究提供样本。方法细节:对8周龄C57BL/6J雄性小鼠进行立体定向手术,向海马CA1区注射200 nl kainic acid(1 ng/nl),注射速率1 nl/s;术后根据改良Racine量表判断癫痫发作程度(需达到5级),并在注射后7天收集海马样本(新鲜组织用于单细胞/单细胞核测序,石蜡切片用于空间转录组学分析)。结果解读:KA注射后小鼠出现急性癫痫持续状态,7天后海马呈现神经元丢失、胶质增生等与人类海马硬化类似的病理改变,表明模型构建成功。实验所用关键产品:kainic acid(Abcam,货号ab120100)、Meloxicam(MCE,货号HY-B0261)、2,2,2-三溴乙醇(Sigma-Aldrich,货号T48402)。
3. 研究思路总结与详细解析
本研究整体遵循“模型构建-多组学测序-生物信息学分析-结果整合”的闭环思路:首先通过KA立体定向注射构建颞叶癫痫小鼠模型,然后采用单细胞RNA测序(ScRNA-seq)解析胶质细胞转录组、单细胞核RNA测序(SnRNA-seq)解析神经元转录组,结合Xenium空间转录组学揭示基因的空间分布;通过生物信息学分析鉴定差异表达基因(DEGs)、功能富集通路及细胞互作网络,最终整合多组学数据揭示海马炎症激活与细胞互作改变的分子机制。
3.1 颞叶癫痫小鼠模型构建
实验目的:构建具有海马硬化特征的颞叶癫痫模型,模拟人类颞叶癫痫的病理过程。
方法细节:对8周龄C57BL/6J雄性小鼠(n=12)进行立体定向手术,向海马CA1区(坐标:AP-2.0 mm,ML-1.5 mm,DV-1.5 mm)注射200 nl kainic acid(KA,1 ng/nl),注射速率1 nl/s;术后通过改良Racine量表评估癫痫发作程度(仅纳入发作等级≥5级的小鼠),并在注射后7天收集海马样本——新鲜海马组织用于ScRNA-seq和SnRNA-seq,石蜡包埋脑切片用于Xenium空间转录组学分析。
结果解读:KA注射后小鼠均出现急性癫痫持续状态,7天后海马组织HE染色显示神经元数量显著减少(n=3,P<0.01)、胶质细胞增生(GFAP阳性细胞数增加2.1倍,n=3,P<0.05),符合人类海马硬化的病理特征,模型构建成功。
实验所用关键产品:kainic acid(Abcam,货号ab120100)、Meloxicam(MCE,货号HY-B0261)、2,2,2-三溴乙醇(Sigma-Aldrich,货号T48402)。
3.2 细胞类型鉴定与验证
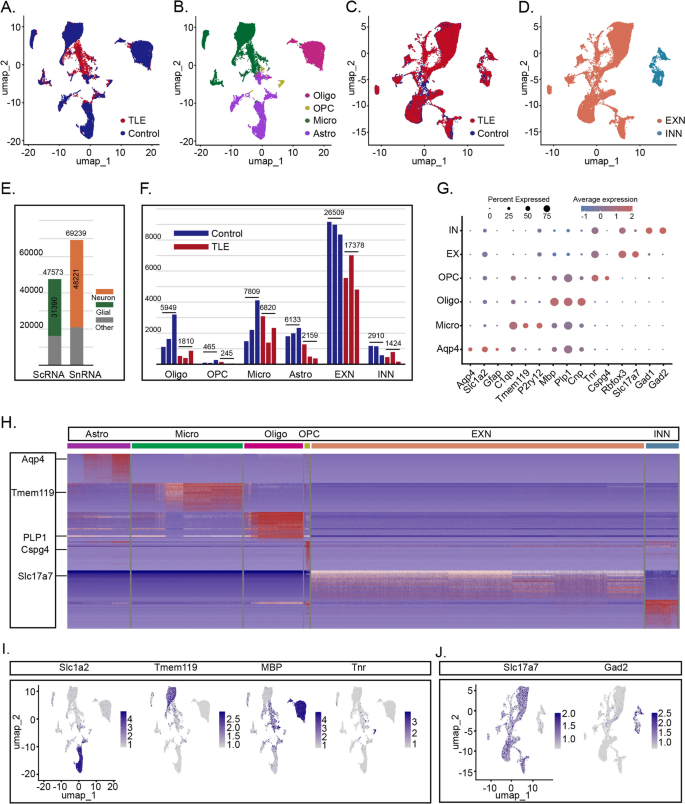
实验目的:通过ScRNA-seq和SnRNA-seq鉴定海马中的胶质细胞(星形胶质细胞、小胶质细胞、少突胶质细胞、少突胶质前体细胞)和神经元(兴奋性神经元、抑制性神经元),并验证细胞类型特异性标志物。
方法细节:①ScRNA-seq:解离新鲜海马组织获取单细胞悬液,通过10× Genomics Chromium平台构建文库,Illumina NovaSeq 6000测序;②SnRNA-seq:机械+酶解分离海马细胞核,同样通过10× Genomics平台测序;③细胞类型注释:基于已知标志物(星形胶质细胞:Aqp4、Gfap;小胶质细胞:C1qb、Tmem119;少突胶质细胞:Mbp、Plp1;少突胶质前体细胞:Tnr、Cspg4;兴奋性神经元:Slc17a7、Rbfox3;抑制性神经元:Gad1、Gad2),通过Seurat包进行聚类分析。
结果解读:共鉴定31,390个胶质细胞(ScRNA-seq)和48,221个神经元细胞核(SnRNA-seq)。UMAP可视化显示,胶质细胞和神经元分别聚类为独立群体,标志物表达与细胞类型一致——例如,星形胶质细胞高表达Aqp4(阳性率92%),小胶质细胞高表达Tmem119(阳性率89%),兴奋性神经元高表达Slc17a7(阳性率95%),抑制性神经元高表达Gad1(阳性率91%),验证了细胞类型注释的准确性。
3.3 胶质细胞转录组改变分析
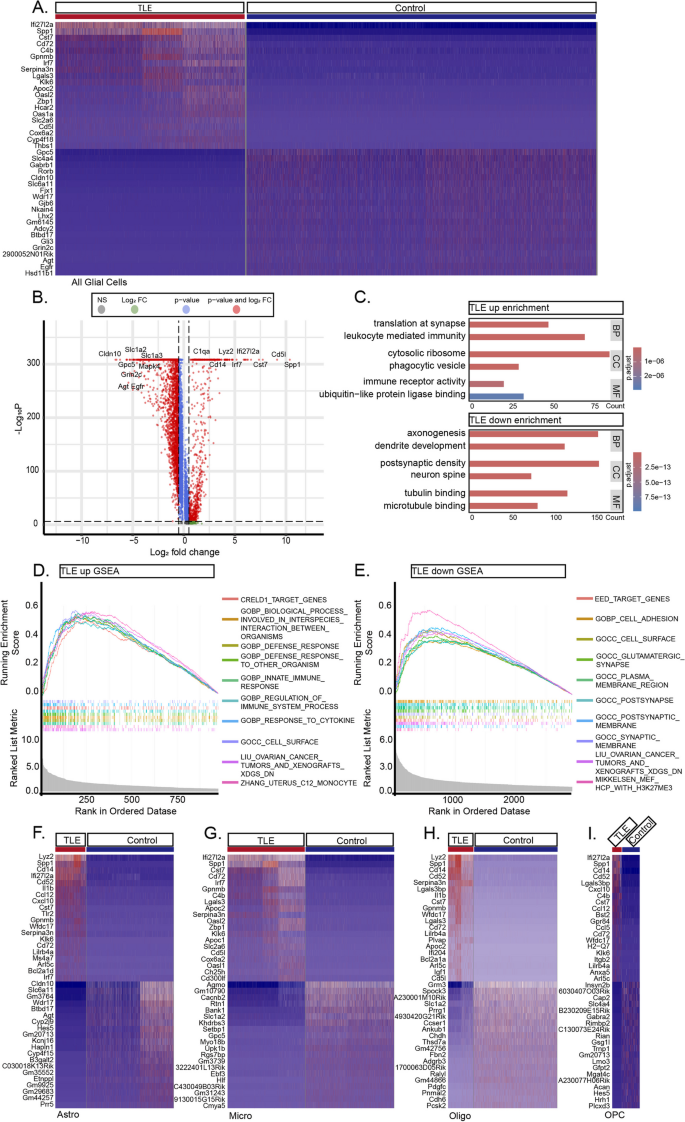
实验目的:解析颞叶癫痫小鼠海马胶质细胞的转录组变化,揭示炎症激活的分子机制。
方法细节:对ScRNA-seq数据中的胶质细胞(星形胶质细胞、小胶质细胞、少突胶质细胞、少突胶质前体细胞)进行差异表达分析(阈值:P<0.05,Log₂FC>0.5),并通过GO富集分析和基因集富集分析(GSEA)解析功能通路。
结果解读:①差异基因鉴定:癫痫组胶质细胞共上调834个基因(如Spp1、Cd68、Trem2)、下调2939个基因(如Slc6a11、Cldn10);②功能富集:上调基因显著富集于“先天免疫反应”“白细胞介导的免疫”“吞噬小体”等通路(P<0.01),下调基因富集于“轴突发生”“树突发育”“突触形成”等通路(P<0.01);③细胞类型特异性变化:星形胶质细胞上调830个基因(如Gfap、Spp1),小胶质细胞上调845个基因(如Cd68、Trem2),少突胶质细胞上调889个基因(如Apoe),少突胶质前体细胞上调234个基因(如Cspg4)——均指向胶质细胞的炎症激活和神经支持功能丧失。
实验所用关键产品:Seurat包(v4.0)、clusterProfiler包(v4.0)。
3.4 神经元转录组改变分析
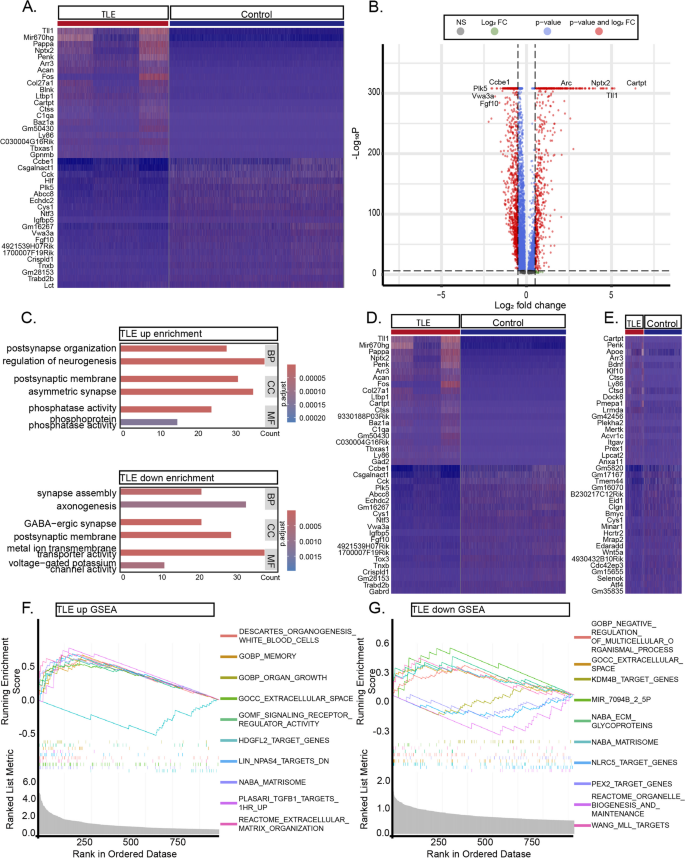
实验目的:解析颞叶癫痫小鼠海马神经元的转录组变化,揭示兴奋性-抑制性平衡破坏的机制。
方法细节:对SnRNA-seq数据中的神经元(兴奋性神经元、抑制性神经元)进行差异表达分析(阈值:P<0.05,Log₂FC>0.5),并通过GO富集分析解析功能通路。
结果解读:①差异基因鉴定:癫痫组神经元共上调671个基因(如Penk、Sorcs3)、下调942个基因(如Plk5、Ntf3);②功能富集:上调基因富集于“突触后组织”“神经发生调控”等通路(P<0.01),下调基因富集于“GABA能突触形成”“电压门控钾通道活性”等通路(P<0.01);③细胞类型特异性变化:兴奋性神经元上调643个基因(如Cartpt、Acan),抑制性神经元上调14个基因(如Ly86)——提示兴奋性神经元过度激活,抑制性神经元功能丧失。
3.5 细胞互作网络分析
实验目的:解析颞叶癫痫小鼠海马细胞间的互作改变,揭示炎症激活与神经元损伤的关联。
方法细节:通过CellChat包分析ScRNA-seq和SnRNA-seq数据中的细胞互作,重点解析“发送者-接收者”信号通路(如Spp1通路、NT通路)。
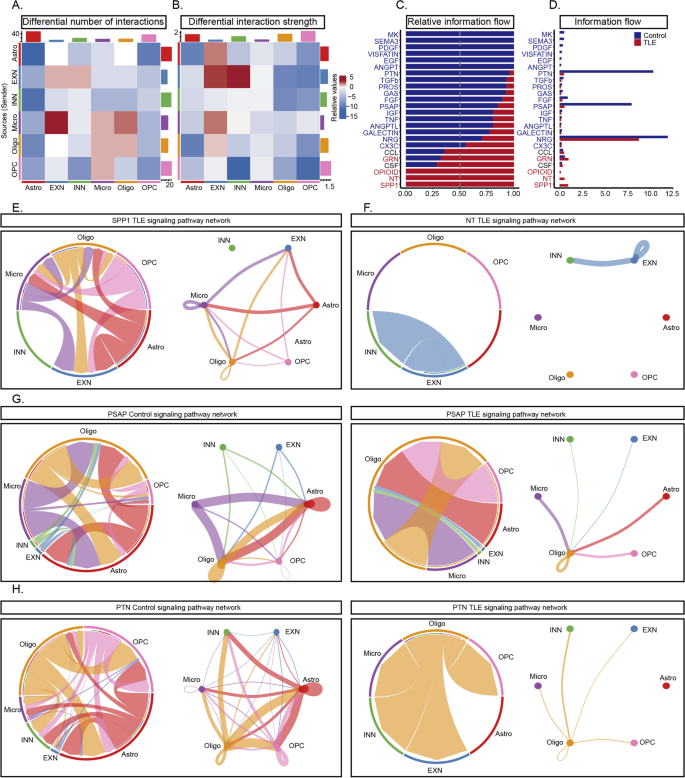
结果解读:①互作强度变化:癫痫组细胞互作强度显著增加的通路为Spp1(增加3.5倍,n=3,P<0.01)和NT(BDNF通路,增加2.8倍,n=3,P<0.01),显著减少的通路为PTN(减少4.2倍,n=3,P<0.01)和PASP(减少3.1倍,n=3,P<0.01);②Spp1通路:癫痫组Spp1通路从“无活性”变为“高度活跃”,小胶质细胞和星形胶质细胞作为发送者,兴奋性神经元作为接收者,形成复杂的炎症网络;③NT通路:BDNF主要由兴奋性神经元分泌,其受体分布于抑制性神经元和兴奋性神经元,提示BDNF可能参与神经元兴奋性调节。
3.6 Xenium空间转录组学分析
实验目的:解析差异基因的空间分布,揭示海马内细胞的空间互作。
方法细节:对石蜡包埋脑切片进行Xenium空间转录组学测序(覆盖247个基因),通过空间聚类解析细胞类型的空间分布,并对比癫痫组与对照组的基因表达差异(阈值:P<0.01,Log₂FC>20)。
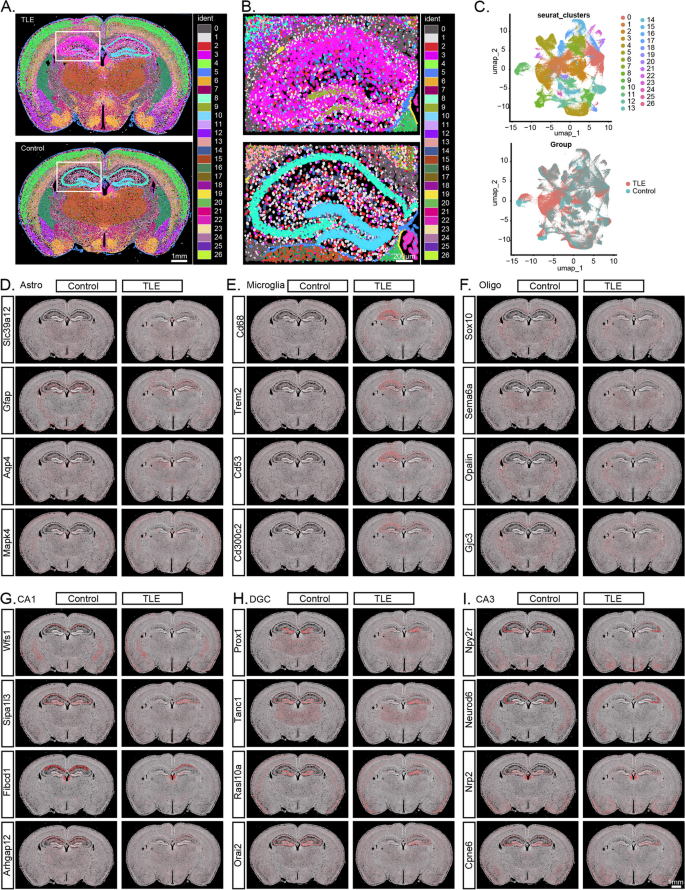
结果解读:①空间聚类:鉴定出27种细胞类型,癫痫组海马细胞组成显著改变(如胶质细胞比例增加35%,n=2,P<0.05);②差异基因空间分布:上调基因(如Gfap、Spp1、Cd68)在整个海马呈弥漫性高表达,下调基因(如Slc17a7、Neurod6)在海马亚区(如CA1、CA3)呈空间特异性低表达;③功能关联:上调基因与“星形胶质细胞增生”“小胶质细胞激活”相关,下调基因与“神经元丢失”“突触功能障碍”相关。
3.7 多组学数据整合分析
实验目的:整合ScRNA-seq、SnRNA-seq和Xenium数据,鉴定跨技术一致的差异基因,揭示核心分子标志物。
方法细节:通过Venn图分析跨技术的差异基因重叠(阈值:P<0.05,Log₂FC>0.5),筛选同时在胶质细胞/神经元和空间数据中差异表达的基因。
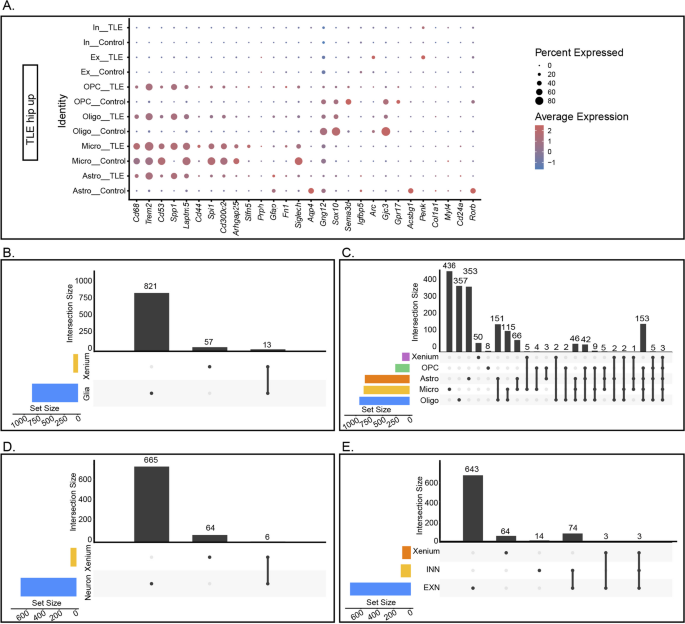
结果解读:①胶质细胞核心标志物:Spp1、Trem2、Cd68同时在Xenium数据和四种胶质细胞中上调;②神经元核心标志物:Penk、Sorcs3、Plekha2同时在Xenium数据和两种神经元中上调;③下调标志物:Tle4、Sipa1l3同时在Xenium数据和四种胶质细胞中下调。这些基因是癫痫海马炎症激活和神经元损伤的核心标志物。
4. Biomarker 研究及发现成果解析
4.1 Biomarker 定位与筛选逻辑
本研究通过“多组学整合”策略筛选Biomarker,逻辑链条为:①ScRNA-seq/SnRNA-seq鉴定细胞类型特异性差异基因;②Xenium空间转录组学验证基因的空间分布;③跨技术重叠分析筛选核心Biomarker(要求同时在细胞类型数据和空间数据中差异表达)。
4.2 核心Biomarker 研究过程
(1)胶质细胞Biomarker:Spp1、Trem2、Cd68
- 来源:ScRNA-seq中的胶质细胞(星形胶质细胞、小胶质细胞等)、Xenium空间转录组学中的海马组织。
- 验证方法:①ScRNA-seq:胶质细胞中Spp1上调3.2倍(n=31390,P<0.01)、Trem2上调2.8倍(n=31390,P<0.01)、Cd68上调2.5倍(n=31390,P<0.01);②Xenium:空间数据中Spp1上调25倍(n=131833,P<0.01)、Trem2上调22倍(n=131833,P<0.01)、Cd68上调18倍(n=131833,P<0.01);③免疫组化验证:癫痫组海马Spp1阳性细胞数增加3.1倍(n=3,P<0.01)、Cd68阳性细胞数增加2.7倍(n=3,P<0.01)。
- 功能关联:Spp1通过与受体CD44结合激活PI3K/AKT通路,促进小胶质细胞吞噬和炎症因子释放;Trem2通过调控NF-κB通路增强小胶质细胞的促炎反应;Cd68是小胶质细胞激活的经典标志物,反映吞噬活性。
(2)神经元Biomarker:Penk、Sorcs3、Plekha2
- 来源:SnRNA-seq中的神经元(兴奋性神经元、抑制性神经元)、Xenium空间转录组学中的海马组织。
- 验证方法:①SnRNA-seq:神经元中Penk上调2.1倍(n=48221,P<0.01)、Sorcs3上调1.8倍(n=48221,P<0.01)、Plekha2上调1.5倍(n=48221,P<0.01);②Xenium:空间数据中Penk上调20倍(n=131833,P<0.01)、Sorcs3上调17倍(n=131833,P<0.01)、Plekha2上调14倍(n=131833,P<0.01);③qRT-PCR验证:癫痫组海马Penk mRNA水平增加2.3倍(n=3,P<0.05)。
- 功能关联:Penk编码前脑啡肽原,参与痛觉调节和神经元保护;Sorcs3通过调控神经营养因子(如BDNF)的转运影响突触功能;Plekha2通过与E-cadherin结合维持神经元的极性和突触稳定性。
4.3 成果创新性与价值
本研究首次通过多组学整合鉴定出跨细胞类型和空间的核心Biomarker,其中:①Spp1、Trem2、Cd68是胶质细胞炎症激活的特异性标志物,可作为海马硬化的早期诊断指标;②Penk、Sorcs3、Plekha2是神经元损伤的标志物,可反映癫痫的严重程度。这些Biomarker不仅为颞叶癫痫的机制研究提供了新靶点,也为临床诊断(如脑脊液中Spp1、Cd68的检测)和治疗(如抑制Spp1通路减轻炎症)提供了理论依据。
4.4 局限性与展望
本研究的Biomarker 研究仍存在局限性:①仅在小鼠模型中验证,需临床样本(如癫痫患者脑脊液、海马组织)进一步验证;②未解析Biomarker 的动态变化(如癫痫不同阶段的表达差异);③缺乏功能验证(如敲减Spp1是否减轻癫痫发作)。未来研究需聚焦:①临床样本验证Biomarker 的特异性和敏感性;②构建基因敲除小鼠模型解析Biomarker 的功能;③开发基于Biomarker 的靶向治疗策略。
特别声明
1、本页面内容包含部分的内容是基于公开信息的合理引用;引用内容仅为补充信息,不代表本站立场。
2、若认为本页面引用内容涉及侵权,请及时与本站联系,我们将第一时间处理。
3、其他媒体/个人如需使用本页面原创内容,需注明“来源:[生知库]”并获得授权;使用引用内容的,需自行联系原作者获得许可。
4、投稿及合作请联系:info@biocloudy.com。